Statistical issues in general (Part I)
March 27, 2023
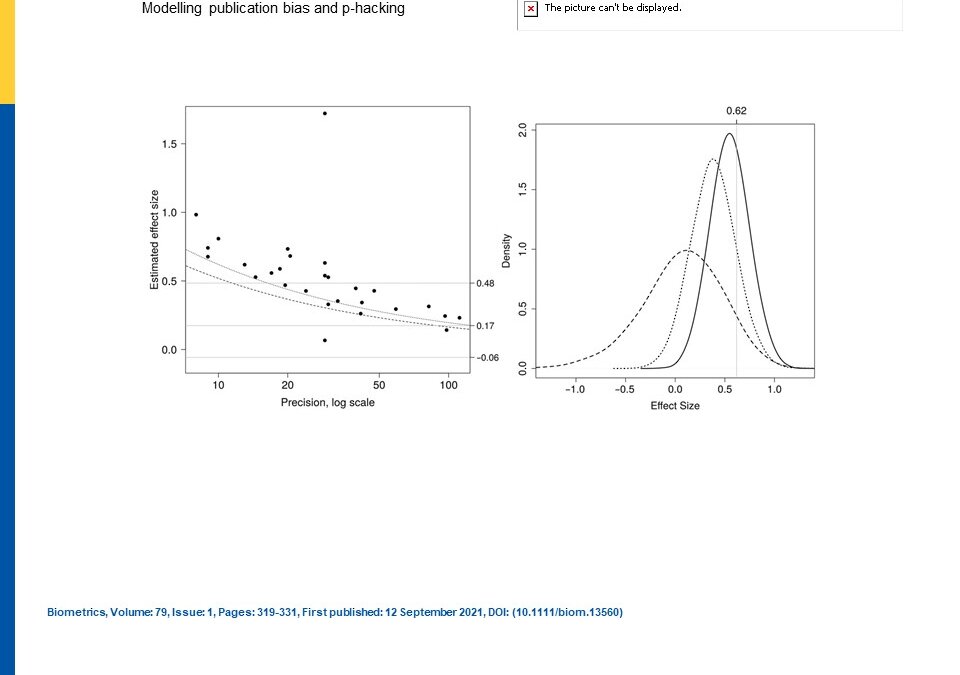
In an article that first appeared in 2021 in Biometrics but is now finally open access in the same journal , the authors, Moss and Bin discuss how publication bias and p-hacking have had strong effects on scientific publication and strongly affect the meta-analyses. P-hacking can be “questionable research practices” or “fishing for significance”, where authors would have manipulated their study results to obtain statistical significance.
The authors first demonstrated their publication bias model where the underlying density of the study-specific parameter is transformed by a selection probability into a publication bias model (Eq 1). This model had two forms with fixed effects or random effects. The authors then demonstrated their p-hacking model, a selection model, based on a normal distribution for each of tractability in showing how the original density with the main parameter of inference is transformed into a p-hacked density (Eq 8), a mixture model, by the researcher tweaking results to confine the results to be drawn from a specific distribution of alpha so that a particular significance level can be reached. This is then a p-hacking model.
In their simulations of meta-analytic studies with varying sample sizes, variability, and various effect sizes and standard deviations for these distributions as well as assumed probabilities of acceptances of papers and p-hacking probabilities, the authors used Bayesian models instead of Frequentist. They found when heterogeneity was reasonable that both publication model and p-hacking model both perform well, but if the effect size is small or the study is small, then the publication model performs worse. They also tried both models on two other example datasets that were meta-analyses. They found that both their models indicated selection bias in these studies as compared to an uncorrected model.
In conclusion, the authors said their p-hacking model worked very well in the simulations but in practice, it was more difficult to discern the differences between this and the publication bias model. One thing they had not done for either model was to allow the parameter of interest to depend upon covariates. This might help but since the p-hacking model is dependent upon the sample size then this could cause more problems with smaller n. Therefore, it would have appeared at the end that the authors suggest running both models and then determining which has the best fit to use. It would seem that trying to emulate the trajectory of p-hacking can be very difficult given the unknown circumstances of how many times or ways that the hacker tries to force a study to obtain statistical significance. They did mention keeping track of the measure of heterogeneity and moving forward, this would always be important to understand the variability amongst studies and also potentially the differences in bias amongst studies, some of which could be induced by p-hacking.
Written by,
Usha Govindarajulu
Keywords: p-values, p-hacking, publication bias, statistical significance
References
Moss J and De Bin R. (2021). “Modelling publication bias and p-hacking’ Biometrics. 79(1): 319-331.
https://doi.org/10.1111/biom.13560
https://onlinelibrary.wiley.com/cms/asset/9df63cf7-505d-440a-a9f8-c39c6bc563c3/biom13560-fig-0001-m.jpg