April 8, 2026
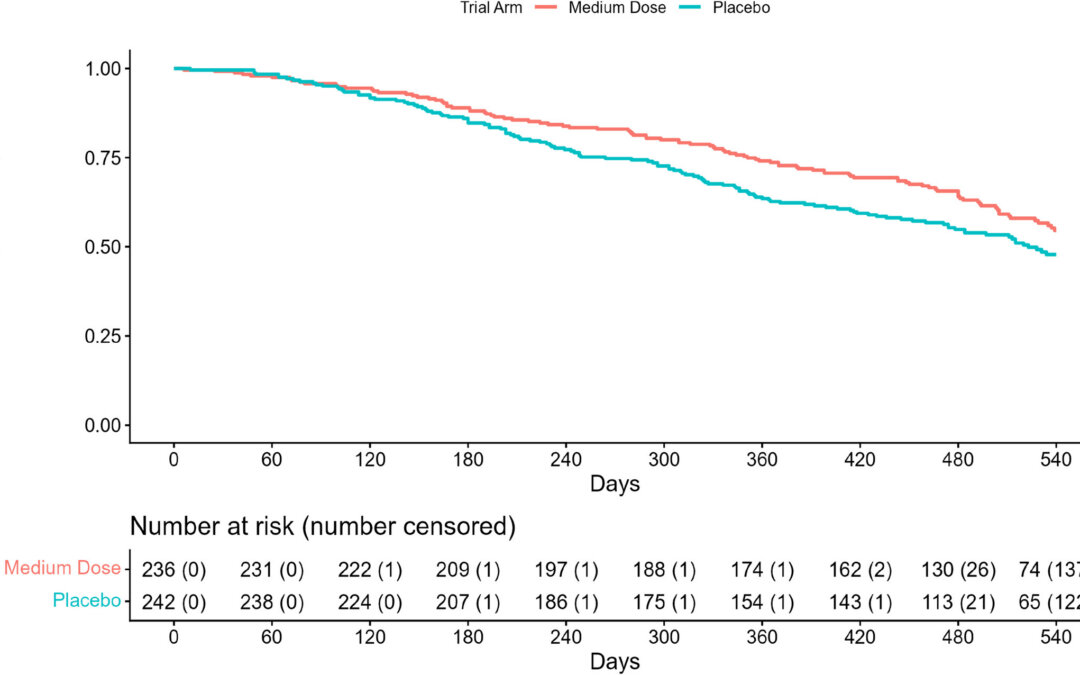
The Restricted Mean Survival Time (RMST) is a metric to compare survival between two treatment groups without relying on proportional hazards, especially that hazard ratios are constant over time. For covariate adjustment, Andersen et al’s (2004) method involves constructing pseudo-values by recalculating the RMST after systematically removing each individual’s data (a “leave-one-out” procedure). The set of pseudo-values, which are not individual RMSTs, can be treated as the outcome variable in a regression model. By regressing the pseudo-values on treatment and other baseline covariates, one can obtain covariate-adjusted estimates of differences in RMST using standard regression techniques. Tian et al (2018) treated the restricted event time, the minimum of the event time and the prespecified truncation time, as the outcome used inverse probability of censoring weights. This way the RMST was directly modeled as a function of covariates, allowing for individual prediction of the RMST. However, these two and other approaches rely on model-based assumptions (i.e. parametric models for hazard or survival or directly relating RMST to covariates or correctly specified propensity score models).
This article by Krajewski and Koch (2026) then introduced a method with minimal assumptions for covariates adjustment to compare survival estimates in randomization-based analysis of covariance (RB-ANCOVA). In this approach, weighted least squares regression is used to produce covariate-adjusted estimates of treatment differences in the outcomes by constraining the differences in covariate means across treatment groups to zero on the basis of randomization. Their approach an accommodate multiple time intervals and therefore estimation of treatment differences at several time points or intervals. In their framework, the RB-ANCOVA is applied to differences in RMSTs along with the weighted least squares estimate.
In a real data analysis, they compared their method to Tian et al’s (2018) ANCOVA method and Andersen et al’s (2004) pseudo value method. They said that their method provided similar estimates and confidence intervals as these methods, however, their method did not require model based assumptions. All their method requires is valid randomization and sufficient sample size. They also reported that in their simulations as well that there was little sensitivity to choice of specific number of intervals, K.
They have given practical suggestions for implementation (sample size, event count, and number of intervals). Their future work would focus on accounting for strata. Some of things that the article did not discuss was the choice of tau and also the fact that they are focus on an ANCOVA, which would suggest normality assumptions
Written by,
Usha Govindarajulu
Keywords: survival, RMST, randomization, confidence intervals, covariate adjustment
References:
K. Andersen, M. G. Hansen, and J. P. Klein, “Regression Analysis of Restricted Mean Survival Time Based on Pseudo-Observations,” Lifetime Data Analysis10, no. 4 (2004): 335–350.
T.Krajewski and G.Koch, “Randomization-Based Covariance Analysis for Confidence Intervals of Treatment Comparisons Based on Restricted Mean Survival Time With Categorized Time-to-Event Data,” Statistics in Medicine45, no. 8-9 (2026): e70422, https://doi.org/10.1002/sim.70422.
Tian, H. Fu, S. J. Ruberg, H. Uno, and L. J. Wei, “Efficiency of Two Sample Tests via the Restricted Mean Survival Time for Analyzing Event Time Observations,” Biometrics74, no. 2 (2018): 694–702.
https://onlinelibrary.wiley.com/cms/asset/919ba05f-7b80-409f-9b6e-f257086d4f86/sim70422-fig-0001-m.jpg